dg/advection_diffusion_2D.py¶
Description
Static advection-diffusion equation in 2D solved by discontinous Galerkin method.
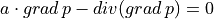
Based on
- Antonietti, P., & Quarteroni, A. (2013). Numerical performance of discontinuous
and stabilized continuous Galerkin methods for convection-diffusion problems.
Usage Examples¶
Run:
sfepy-run sfepy/examples/dg/advection_diffusion_2D.py
Results are saved to output/dg/advection_diffusion_2D folder by default as `
.msh` files, the best way to view them is through GMSH (http://gmsh.info/)
version 4.6 or newer. Start GMSH and use File | Open menu or Crtl + O
shortcut, navigate to the output folder, select all .msh files and hit Open,
all files should load as one item in Post-processing named p_cell_nodes.
GMSH is capable of rendering high order approximations in individual elements,
to modify fidelity of rendering, double click the displayed mesh, quick options
menu should pop up, click on All view options.... This brings up the Options
window with View [0] selected in left column. Under the tab General
ensure that Adapt visualization grid is ticked, then you can adjust
Maximum recursion depth and `Target visualization error to tune
the visualization. To see visualization elements (as opposed to mesh elements)
go to Visibility tab and tick Draw element outlines, this option is also
available from quick options menu as View element outlines or under shortcut
Alt+E. In the quick options menu, you can also modify normal raise by
clicking View Normal Raise to see solution rendered as surface above the
mesh. Note that for triangular meshes normal raise -1 produces expected raise
above the mesh. This is due to the opposite orientation of the reference
elements in GMSH and Sfepy and might get patched in the future.
r"""
Static advection-diffusion equation in 2D solved by discontinous Galerkin method.
.. math:: a \cdot grad\, p - div(grad\,p) = 0
Based on
Antonietti, P., & Quarteroni, A. (2013). Numerical performance of discontinuous
and stabilized continuous Galerkin methods for convection-diffusion problems.
Usage Examples
--------------
Run::
sfepy-run sfepy/examples/dg/advection_diffusion_2D.py
Results are saved to output/dg/advection_diffusion_2D folder by default as `
`.msh`` files, the best way to view them is through GMSH (http://gmsh.info/)
version 4.6 or newer. Start GMSH and use ``File | Open`` menu or Crtl + O
shortcut, navigate to the output folder, select all ``.msh`` files and hit Open,
all files should load as one item in Post-processing named p_cell_nodes.
GMSH is capable of rendering high order approximations in individual elements,
to modify fidelity of rendering, double click the displayed mesh, quick options
menu should pop up, click on ``All view options...``. This brings up the Options
window with ``View [0]`` selected in left column. Under the tab ``General``
ensure that ``Adapt visualization grid`` is ticked, then you can adjust
``Maximum recursion depth`` and ```Target visualization error`` to tune
the visualization. To see visualization elements (as opposed to mesh elements)
go to ``Visibility`` tab and tick ``Draw element outlines``, this option is also
available from quick options menu as ``View element outlines`` or under shortcut
``Alt+E``. In the quick options menu, you can also modify normal raise by
clicking ``View Normal Raise`` to see solution rendered as surface above the
mesh. Note that for triangular meshes normal raise -1 produces expected raise
above the mesh. This is due to the opposite orientation of the reference
elements in GMSH and Sfepy and might get patched in the future.
"""
from sfepy.examples.dg.example_dg_common import *
def define(filename_mesh=None,
approx_order=3,
adflux=0,
limit=False,
cw=1000,
diffcoef=1,
diffscheme="symmetric",
cfl=None,
dt=None,
):
cfl = None
dt = None
functions = {}
def local_register_function(fun):
try:
functions.update({fun.__name__: (fun,)})
except AttributeError: # Already a sfepy Function.
fun = fun.function
functions.update({fun.__name__: (fun,)})
return fun
example_name = "advection_diffusion_2D"
dim = 2
if filename_mesh is None:
filename_mesh = get_gen_block_mesh_hook((1., 1.), (20, 20), (.5, .5))
velo = [1., 1.]
angle = 0.0 # - nm.pi / 5
rotm = nm.array([[nm.cos(angle), -nm.sin(angle)],
[nm.sin(angle), nm.cos(angle)]])
velo = nm.sum(rotm.T * nm.array(velo), axis=-1)[:, None]
regions = {
'Omega' : 'all',
'left' : ('vertices in x == 0', 'edge'),
'right': ('vertices in x == 1', 'edge'),
'top' : ('vertices in y == 1', 'edge'),
'bottom': ('vertices in y == 0', 'edge')
}
fields = {
'f': ('real', 'scalar', 'Omega', str(approx_order) + 'd', 'DG', 'legendre')
}
variables = {
'p': ('unknown field', 'f', 0),
'v': ('test field', 'f', 'p'),
}
integrals = {
'i': 2 * approx_order,
}
@local_register_function
def bc_funs(ts, coors, bc, problem):
# return 2*coors[..., 1]
t = ts.dt*ts.step
x_1 = coors[..., 0]
x_2 = coors[..., 1]
res = nm.zeros(nm.shape(x_1))
sin = nm.sin
cos = nm.cos
exp = nm.exp
pi = nm.pi
if bc.diff == 0:
if "left" in bc.name:
res[:] = 0
elif "right" in bc.name:
res[:] = 0
elif "bottom" in bc.name:
res[:] = 0 #-2*sin(2*pi*x_1)
elif "top" in bc.name:
res[:] = 0
elif bc.diff == 1:
if "left" in bc.name:
res = nm.stack((-2*pi*(x_2**2 - x_2),
res),
axis=-2)
elif "right" in bc.name:
res = nm.stack((-2*pi*(x_2**2 - x_2), res,),
axis=-2)
elif "bot" in bc.name:
res = nm.stack((res,
sin(2*pi*x_1)),
axis=-2)
elif "top" in bc.name:
res = nm.stack((res,
-sin(2*pi*x_1)),
axis=-2)
return res
@local_register_function
def source_fun(ts, coors, mode="qp", **kwargs):
# t = ts.dt * ts.step
eps = diffcoef
sin = nm.sin
cos = nm.cos
exp = nm.exp
sqrt = nm.sqrt
pi = nm.pi
if mode == "qp":
x_1 = coors[..., 0]
x_2 = coors[..., 1]
res = -2*pi*(x_2**2 - x_2)*cos(2*pi*x_1)\
- 2*(2*pi**2*(x_2**2 - x_2)*sin(2*pi*x_1) - sin(2*pi*x_1))*eps\
- (2*x_2 - 1)*sin(2*pi*x_1)
return {"val": res[..., None, None]}
def analytic_sol(coors, t):
x_1 = coors[..., 0]
x_2 = coors[..., 1]
sin = nm.sin
pi = nm.pi
res = -(x_2 ** 2 - x_2) * sin(2 * pi * x_1)
return res
@local_register_function
def sol_fun(ts, coors, mode="qp", **kwargs):
t = ts.time
if mode == "qp":
return {"p": analytic_sol(coors, t)[..., None, None]}
dgebcs = {
'u_left' : ('left', {'p.all': "bc_funs", 'grad.p.all' : "bc_funs"}),
'u_top' : ('top', {'p.all': "bc_funs", 'grad.p.all' : "bc_funs"}),
'u_bot' : ('bottom', {'p.all': "bc_funs", 'grad.p.all' : "bc_funs"}),
'u_right': ('right', {'p.all': "bc_funs", 'grad.p.all' : "bc_funs"}),
}
materials = {
'a' : ({'val': [velo], '.flux': adflux},),
'D' : ({'val': [diffcoef], '.cw': cw},),
'g' : 'source_fun'
}
equations = {
'balance': """
- dw_s_dot_mgrad_s.i.Omega(a.val, p, v)
+ dw_dg_advect_laxfrie_flux.i.Omega(a.flux, a.val, v, p)
"""
+
" + dw_laplace.i.Omega(D.val, v, p) " +
diffusion_schemes_implicit[diffscheme] +
" + dw_dg_interior_penalty.i.Omega(D.val, D.cw, v, p)" +
" - dw_volume_lvf.i.Omega(g.val, v)" +
"= 0"
}
solver_0 = {
'name' : 'ls',
'kind' : 'ls.scipy_direct',
}
solver_1 = {
'name' : 'newton',
'kind' : 'nls.newton',
'i_max' : 5,
'eps_a' : 1e-8,
'eps_r' : 1.0,
'macheps' : 1e-16,
'lin_red' : 1e-2, # Linear system error < (eps_a * lin_red).
'ls_red' : 0.1,
'ls_red_warp' : 0.001,
'ls_on' : 0.99999,
'ls_min' : 1e-5,
'check' : 0,
'delta' : 1e-6,
}
options = {
'nls' : 'newton',
'ls' : 'ls',
'output_dir' : 'output/dg/' + example_name,
'output_format' : 'msh',
'file_format' : 'gmsh-dg'
}
return locals()