multi_physics/biot_npbc_lagrange.py¶
Description
Biot problem - deformable porous medium with the no-penetration boundary condition on a boundary region enforced using Lagrange multipliers.
The non-penetration condition is enforced weakly using the Lagrange
multiplier 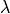 . There is also a rigid body movement
constraint imposed on the
. There is also a rigid body movement
constraint imposed on the 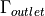 region using the
linear combination boundary conditions.
region using the
linear combination boundary conditions.
Find 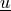 ,
, 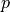 and
and 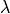 such that:
such that:
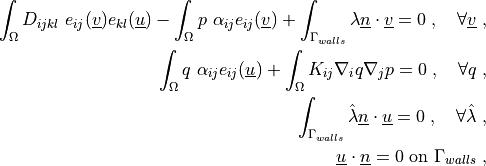
where
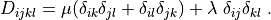
r"""
Biot problem - deformable porous medium with the no-penetration boundary
condition on a boundary region enforced using Lagrange multipliers.
The non-penetration condition is enforced weakly using the Lagrange
multiplier :math:`\lambda`. There is also a rigid body movement
constraint imposed on the :math:`\Gamma_{outlet}` region using the
linear combination boundary conditions.
Find :math:`\ul{u}`, :math:`p` and :math:`\lambda` such that:
.. math::
\int_{\Omega} D_{ijkl}\ e_{ij}(\ul{v}) e_{kl}(\ul{u})
- \int_{\Omega} p\ \alpha_{ij} e_{ij}(\ul{v})
+ \int_{\Gamma_{walls}} \lambda \ul{n} \cdot \ul{v}
= 0
\;, \quad \forall \ul{v} \;,
\int_{\Omega} q\ \alpha_{ij} e_{ij}(\ul{u})
+ \int_{\Omega} K_{ij} \nabla_i q \nabla_j p
= 0
\;, \quad \forall q \;,
\int_{\Gamma_{walls}} \hat\lambda \ul{n} \cdot \ul{u}
= 0
\;, \quad \forall \hat\lambda \;,
\ul{u} \cdot \ul{n} = 0 \mbox{ on } \Gamma_{walls} \;,
where
.. math::
D_{ijkl} = \mu (\delta_{ik} \delta_{jl}+\delta_{il} \delta_{jk}) +
\lambda \ \delta_{ij} \delta_{kl}
\;.
"""
from sfepy.examples.multi_physics.biot_npbc import (cinc_simple,
define_regions, get_pars)
def define():
from sfepy import data_dir
filename = data_dir + '/meshes/3d/cylinder.mesh'
output_dir = 'output'
return define_input(filename, output_dir)
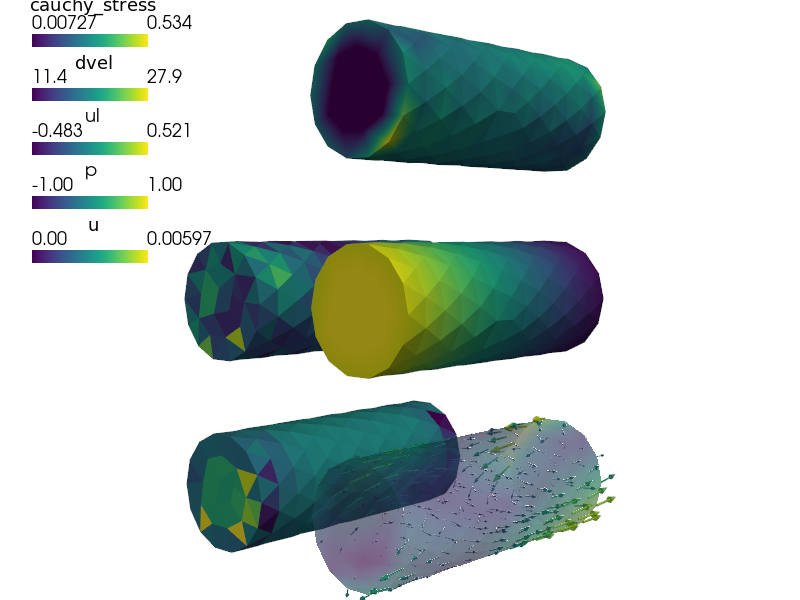
def post_process(out, pb, state, extend=False):
from sfepy.base.base import Struct
dvel = pb.evaluate('ev_diffusion_velocity.2.Omega( m.K, p )',
mode='el_avg')
out['dvel'] = Struct(name='output_data', var_name='p',
mode='cell', data=dvel, dofs=None)
stress = pb.evaluate('ev_cauchy_stress.2.Omega( m.D, u )',
mode='el_avg')
out['cauchy_stress'] = Struct(name='output_data', var_name='u',
mode='cell', data=stress, dofs=None)
return out
def define_input(filename, output_dir):
filename_mesh = filename
options = {
'output_dir' : output_dir,
'output_format' : 'vtk',
'post_process_hook' : 'post_process',
'ls' : 'ls',
'nls' : 'newton',
}
functions = {
'cinc_simple0' : (lambda coors, domain:
cinc_simple(coors, 0),),
'cinc_simple1' : (lambda coors, domain:
cinc_simple(coors, 1),),
'cinc_simple2' : (lambda coors, domain:
cinc_simple(coors, 2),),
'get_pars' : (lambda ts, coors, mode=None, **kwargs:
get_pars(ts, coors, mode,
output_dir=output_dir, **kwargs),),
}
regions, dim = define_regions(filename_mesh)
fields = {
'displacement': ('real', 'vector', 'Omega', 1),
'pressure': ('real', 'scalar', 'Omega', 1),
'multiplier': ('real', 'scalar', 'Walls', 1),
}
variables = {
'u' : ('unknown field', 'displacement', 0),
'v' : ('test field', 'displacement', 'u'),
'p' : ('unknown field', 'pressure', 1),
'q' : ('test field', 'pressure', 'p'),
'ul' : ('unknown field', 'multiplier', 2),
'vl' : ('test field', 'multiplier', 'ul'),
}
ebcs = {
'inlet' : ('Inlet', {'p.0' : 1.0, 'u.all' : 0.0}),
'outlet' : ('Outlet', {'p.0' : -1.0}),
}
lcbcs = {
'rigid' : ('Outlet', {'u.all' : None}, None, 'rigid'),
}
materials = {
'm' : 'get_pars',
}
equations = {
'eq_1' :
"""dw_lin_elastic.2.Omega( m.D, v, u )
- dw_biot.2.Omega( m.alpha, v, p )
+ dw_non_penetration.2.Walls( v, ul )
= 0""",
'eq_2' :
"""dw_biot.2.Omega( m.alpha, u, q )
+ dw_diffusion.2.Omega( m.K, q, p )
= 0""",
'eq_3' :
"""dw_non_penetration.2.Walls( u, vl )
= 0""",
}
solvers = {
'ls' : ('ls.scipy_direct', {}),
'newton' : ('nls.newton', {}),
}
return locals()