linear_elasticity/elastic2D_axisymmetric.py¶
Description
Axisymmetric Linear Elasticity (2D formulation).
This example demonstrates a two-dimensional axisymmetric formulation of the three-dimensional static linear elasticity equations. The formulation is derived under the standard axisymmetry assumptions and is equivalent to the original three-dimensional problem.
Three-dimensional linear elasticity¶
Let 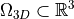 denote an elastic body. The static
equilibrium equations read
denote an elastic body. The static
equilibrium equations read
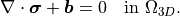
In the absence of body forces (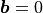 ),
),
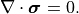
The material is assumed to be linear, isotropic, and homogeneous, with the constitutive relation
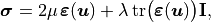
where the infinitesimal strain tensor is defined as
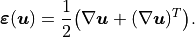
Axisymmetric assumptions¶
Cylindrical coordinates 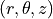 are introduced. The following
assumptions are made:
are introduced. The following
assumptions are made:
all fields are independent of
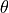 ,
,the circumferential displacement vanishes:
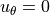 .
.
The displacement field therefore reduces to
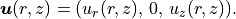
The computational domain is the meridian cross-section
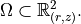
Axisymmetric strong form¶
Radial equilibrium¶
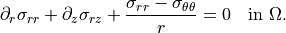
Axial equilibrium¶
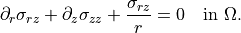
Axisymmetric weak formulation¶
Let 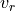 and
and 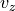 denote admissible test functions. Assuming zero
traction boundary conditions, integration over the circumferential direction
yields the axisymmetric volume element
denote admissible test functions. Assuming zero
traction boundary conditions, integration over the circumferential direction
yields the axisymmetric volume element
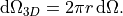
Radial equilibrium (weak form)¶
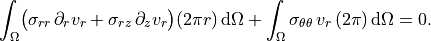
Axial equilibrium (weak form)¶
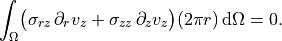
Stress–strain relations in axisymmetry¶
The nonzero stress components in axisymmetry are
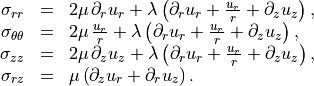
Compact bilinear form¶
Radial test function¶
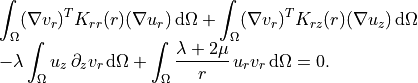
Axial test function¶
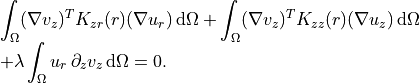
Axisymmetric material matrices¶
![K_{rr}(r) = r
\left[
\begin{array}{cc}
\lambda + 2\mu & 0 \\
0 & \mu
\end{array}
\right],
\qquad
K_{rz}(r) = r
\left[
\begin{array}{cc}
0 & \lambda \\
\mu & 0
\end{array}
\right].](../_images/math/ee9153062dc02380caaad9ee6c664b0103a6ff57.png)
![K_{zr}(r) = r
\left[
\begin{array}{cc}
0 & \mu \\
\lambda & 0
\end{array}
\right],
\qquad
K_{zz}(r) = r
\left[
\begin{array}{cc}
\mu & 0 \\
0 & \lambda + 2\mu
\end{array}
\right].](../_images/math/2d61be5ced714ceff6fe2424cdd778e1a14e0d1c.png)
Remark¶
This formulation is mathematically equivalent to the original three-dimensional linear elasticity problem under the stated axisymmetry assumptions. It is directly suitable for two-dimensional finite element discretizations using the axisymmetric weighting.
To run the example, execute one of the following commands:
python3 -m sfepy.scripts.simple ./sfepy/examples/linear_elasticity/elastic2D_axisymmetric.py
sfepy-run sfepy/examples/linear_elasticity/elastic2D_axisymmetric.py

r"""
Axisymmetric Linear Elasticity (2D formulation).
This example demonstrates a two-dimensional axisymmetric formulation of the
three-dimensional static linear elasticity equations. The formulation is derived
under the standard axisymmetry assumptions and is equivalent to the original
three-dimensional problem.
Three-dimensional linear elasticity
-----------------------------------
Let :math:`\Omega_{3D} \subset \mathbb{R}^3` denote an elastic body. The static
equilibrium equations read
.. math::
\nabla \cdot \boldsymbol{\sigma} + \boldsymbol{b} = 0
\quad \text{in } \Omega_{3D}.
In the absence of body forces (:math:`\boldsymbol{b} = 0`),
.. math::
\nabla \cdot \boldsymbol{\sigma} = 0.
The material is assumed to be linear, isotropic, and homogeneous, with the
constitutive relation
.. math::
\boldsymbol{\sigma}
= 2\mu\,\boldsymbol{\varepsilon}(\boldsymbol{u})
+ \lambda\,\mathrm{tr}
\bigl(\boldsymbol{\varepsilon}(\boldsymbol{u})\bigr)\mathbf{I},
where the infinitesimal strain tensor is defined as
.. math::
\boldsymbol{\varepsilon}(\boldsymbol{u})
= \frac{1}{2}\bigl(
\nabla\boldsymbol{u} + (\nabla\boldsymbol{u})^{T}
\bigr).
Axisymmetric assumptions
------------------------
Cylindrical coordinates :math:`(r, \theta, z)` are introduced. The following
assumptions are made:
- all fields are independent of :math:`\theta`,
- the circumferential displacement vanishes: :math:`u_\theta = 0`.
The displacement field therefore reduces to
.. math::
\boldsymbol{u}(r, z) = (u_r(r, z),\, 0,\, u_z(r, z)).
The computational domain is the meridian cross-section
.. math::
\Omega \subset \mathbb{R}^2_{(r, z)}.
Axisymmetric strong form
------------------------
Radial equilibrium
~~~~~~~~~~~~~~~~~~
.. math::
\partial_r \sigma_{rr}
+ \partial_z \sigma_{rz}
+ \frac{\sigma_{rr} - \sigma_{\theta\theta}}{r}
= 0
\quad \text{in } \Omega.
Axial equilibrium
~~~~~~~~~~~~~~~~~
.. math::
\partial_r \sigma_{rz}
+ \partial_z \sigma_{zz}
+ \frac{\sigma_{rz}}{r}
= 0
\quad \text{in } \Omega.
Axisymmetric weak formulation
-----------------------------
Let :math:`v_r` and :math:`v_z` denote admissible test functions. Assuming zero
traction boundary conditions, integration over the circumferential direction
yields the axisymmetric volume element
.. math::
\mathrm{d}\Omega_{3D} = 2\pi r\,\mathrm{d}\Omega.
Radial equilibrium (weak form)
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
.. math::
\int_\Omega
\bigl(
\sigma_{rr}\,\partial_r v_r
+ \sigma_{rz}\,\partial_z v_r
\bigr)(2\pi r)\,\mathrm{d}\Omega
+ \int_\Omega
\sigma_{\theta\theta}\,v_r\,(2\pi)\,\mathrm{d}\Omega
= 0.
Axial equilibrium (weak form)
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
.. math::
\int_\Omega
\bigl(
\sigma_{rz}\,\partial_r v_z
+ \sigma_{zz}\,\partial_z v_z
\bigr)(2\pi r)\,\mathrm{d}\Omega
= 0.
Stress--strain relations in axisymmetry
---------------------------------------
The nonzero stress components in axisymmetry are
.. math::
\begin{array}{rcl}
\sigma_{rr} &=&
2\mu\,\partial_r u_r
+ \lambda\left(\partial_r u_r + \frac{u_r}{r} + \partial_z u_z\right), \\
\sigma_{\theta\theta} &=&
2\mu\,\frac{u_r}{r}
+ \lambda\left(\partial_r u_r + \frac{u_r}{r} + \partial_z u_z\right), \\
\sigma_{zz} &=&
2\mu\,\partial_z u_z
+ \lambda\left(\partial_r u_r + \frac{u_r}{r} + \partial_z u_z\right), \\
\sigma_{rz} &=&
\mu\left(\partial_z u_r + \partial_r u_z\right).
\end{array}
Compact bilinear form
---------------------
Radial test function
~~~~~~~~~~~~~~~~~~~~
.. math::
\begin{array}{l}
\displaystyle
\int_{\Omega} (\nabla v_r)^{T} K_{rr}(r) (\nabla u_r)\,\mathrm{d}\Omega
+ \int_{\Omega} (\nabla v_r)^{T} K_{rz}(r) (\nabla u_z)\,\mathrm{d}\Omega \\
\displaystyle
- \lambda \int_{\Omega} u_z\,\partial_z v_r\,\mathrm{d}\Omega
+ \int_{\Omega} \frac{\lambda + 2\mu}{r}\, u_r v_r\,\mathrm{d}\Omega
= 0.
\end{array}
Axial test function
~~~~~~~~~~~~~~~~~~~
.. math::
\begin{array}{l}
\displaystyle
\int_{\Omega} (\nabla v_z)^{T} K_{zr}(r) (\nabla u_r)\,\mathrm{d}\Omega
+ \int_{\Omega} (\nabla v_z)^{T} K_{zz}(r) (\nabla u_z)\,\mathrm{d}\Omega\\
\displaystyle
+ \lambda \int_{\Omega} u_r\,\partial_z v_z\,\mathrm{d}\Omega
= 0.
\end{array}
Axisymmetric material matrices
------------------------------
.. math::
K_{rr}(r) = r
\left[
\begin{array}{cc}
\lambda + 2\mu & 0 \\
0 & \mu
\end{array}
\right],
\qquad
K_{rz}(r) = r
\left[
\begin{array}{cc}
0 & \lambda \\
\mu & 0
\end{array}
\right].
.. math::
K_{zr}(r) = r
\left[
\begin{array}{cc}
0 & \mu \\
\lambda & 0
\end{array}
\right],
\qquad
K_{zz}(r) = r
\left[
\begin{array}{cc}
\mu & 0 \\
0 & \lambda + 2\mu
\end{array}
\right].
Remark
------
This formulation is mathematically equivalent to the original three-dimensional
linear elasticity problem under the stated axisymmetry assumptions. It is
directly suitable for two-dimensional finite element discretizations using the
axisymmetric weighting.
To run the example, execute one of the following commands::
python3 -m sfepy.scripts.simple ./sfepy/examples/linear_elasticity/elastic2D_axisymmetric.py
sfepy-run sfepy/examples/linear_elasticity/elastic2D_axisymmetric.py
"""
import numpy as np
E = 70e9
nu = 0.3
r_inner, r_outer = 0.1, 0.2
height = 0.3
a, b = 1e-3, 2e-3
def define():
from sfepy.discrete.fem import Mesh
from sfepy.homogenization.utils import define_box_regions
lam = E * nu / ((1 + nu) * (1 - 2 * nu))
mu = E / (2 * (1 + nu))
nr, nz = 9, 11
r = np.linspace(r_inner, r_outer, nr)
z = np.linspace(0.0, height, nz)
nodes = []
for iz in range(nz):
for ir in range(nr):
nodes.append([r[ir], z[iz]])
coors = np.array(nodes, dtype=np.float64)
conn = []
for iz in range(nz - 1):
for ir in range(nr - 1):
n0 = iz * nr + ir
conn.append([n0, n0 + 1, n0 + nr + 1, n0 + nr])
conn = np.array(conn, dtype=np.int32)
filename_mesh = mesh = Mesh.from_data(
"annulus",
coors,
None,
[conn],
[np.zeros(len(conn), dtype=np.int32)],
["2_4"],
)
# Get bounding box for region definition
bbox = mesh.get_bounding_box()
# -----------------------------
# Material coefficient functions
# -----------------------------
def material_rr(ts, coors, mode=None, **kwargs):
if mode == "qp":
rr = coors[:, 0]
K = np.zeros((coors.shape[0], 2, 2), dtype=np.float64)
K[:, 0, 0] = (lam + 2 * mu) * rr
K[:, 1, 1] = mu * rr
return {"K": K}
def material_zz(ts, coors, mode=None, **kwargs):
if mode == "qp":
rr = coors[:, 0]
K = np.zeros((coors.shape[0], 2, 2), dtype=np.float64)
K[:, 0, 0] = mu * rr
K[:, 1, 1] = (lam + 2 * mu) * rr
return {"K": K}
def material_poisson_rr(ts, coors, mode=None, **kwargs):
if mode == "qp":
rr = coors[:, 0]
K = np.zeros((coors.shape[0], 2, 2), dtype=np.float64)
K[:, 0, 1] = mu * rr
K[:, 1, 0] = lam * rr
return {"K": K}
def material_poisson_zz(ts, coors, mode=None, **kwargs):
if mode == "qp":
rr = coors[:, 0]
K = np.zeros((coors.shape[0], 2, 2), dtype=np.float64)
K[:, 0, 1] = lam * rr
K[:, 1, 0] = mu * rr
return {"K": K}
def material_hoop0(ts, coors, mode=None, **kwargs):
if mode == "qp":
rr = coors[:, 0]
rr_safe = np.maximum(rr, 1e-12)
c = (lam + 2 * mu) / rr_safe
return {"c": c.reshape(-1, 1, 1)}
# -----------------------------
# Regions: Use define_box_regions for automatic boundary selection
# -----------------------------
regions = {
"Omega": "all",
}
regions.update(define_box_regions(2, bbox[0], bbox[1]))
materials = {
"m_rr": "material_rr",
"m_zz": "material_zz",
"m_prr": "material_poisson_rr",
"m_pzz": "material_poisson_zz",
"m_hoop_idx": ({'.val' : np.array([[1]])},),
"m_hoop0": "material_hoop0",
}
fields = {
"ur": ("real", 1, "Omega", 1),
"uz": ("real", 1, "Omega", 1),
}
integrals = {"i": 5}
variables = {
"ur": ("unknown field", "ur", 0),
"vr": ("test field", "ur", "ur"),
"uz": ("unknown field", "uz", 1),
"vz": ("test field", "uz", "uz"),
}
# Register both material funcs and region-selection funcs here!
functions = {
"material_rr": (material_rr,),
"material_zz": (material_zz,),
"material_poisson_rr": (material_poisson_rr,),
"material_poisson_zz": (material_poisson_zz,),
"material_hoop0": (material_hoop0,),
# ur = a*r
"bc_ur_left": (lambda ts, coors, **kw: a * r_inner * np.ones(len(coors)),),
"bc_ur_right": (lambda ts, coors, **kw: a * r_outer * np.ones(len(coors)),),
"bc_ur_bottom": (lambda ts, coors, **kw: a * coors[:, 0],),
"bc_ur_top": (lambda ts, coors, **kw: a * coors[:, 0],),
# uz = b*z
"bc_uz_left": (lambda ts, coors, **kw: b * coors[:, 1],),
"bc_uz_right": (lambda ts, coors, **kw: b * coors[:, 1],),
"bc_uz_bottom": (lambda ts, coors, **kw: np.zeros(len(coors)),),
"bc_uz_top": (lambda ts, coors, **kw: b * height * np.ones(len(coors)),),
}
ebcs = {
"fix_ur_left": ("Left", {"ur.0": "bc_ur_left"}),
"fix_ur_right": ("Right", {"ur.0": "bc_ur_right"}),
"fix_ur_bottom": ("Bottom", {"ur.0": "bc_ur_bottom"}),
"fix_ur_top": ("Top", {"ur.0": "bc_ur_top"}),
"fix_uz_left": ("Left", {"uz.0": "bc_uz_left"}),
"fix_uz_right": ("Right", {"uz.0": "bc_uz_right"}),
"fix_uz_bottom": ("Bottom", {"uz.0": "bc_uz_bottom"}),
"fix_uz_top": ("Top", {"uz.0": "bc_uz_top"}),
}
equations = {
"balance_r": r"""
dw_diffusion.i.Omega(m_rr.K, vr, ur)
+ dw_diffusion.i.Omega(m_prr.K, vr, uz)
- dw_s_dot_grad_i_s.i.Omega(m_hoop_idx.val, vr, uz)
+ dw_volume_dot.i.Omega(m_hoop0.c, vr, ur)
= 0
""",
"balance_z": r"""
dw_diffusion.i.Omega(m_zz.K, vz, uz)
+ dw_diffusion.i.Omega(m_pzz.K, vz, ur)
+ dw_s_dot_grad_i_s.i.Omega(m_hoop_idx.val, vz, ur)
= 0
""",
}
solvers = {
"ls": ("ls.scipy_direct", {}),
"newton": ("nls.newton", {"i_max": 1, "eps_a": 1e-12, "eps_r": 1e-12,
"eps_mode": "or", "lin_red": None}),
}
options = {
"nls": "newton",
"ls": "ls",
"post_process_hook": "post_process",
}
return locals()
def post_process(out, problem, variables, extend=False):
ur = variables["ur"]
uz = variables["uz"]
coors = problem.domain.get_mesh_coors()
rr, zz = coors[:, 0], coors[:, 1]
u_r_exact = a * rr
u_z_exact = b * zz
u_r_val = ur().ravel()
u_z_val = uz().ravel()
err_r = np.abs(u_r_val - u_r_exact)
err_z = np.abs(u_z_val - u_z_exact)
err_total = np.sqrt(err_r**2 + err_z**2)
max_err = np.max(err_total)
print("\n" + "=" * 70)
print("AXISYMMETRIC ELASTICITY VIA TERM COMPOSITION")
print("=" * 70)
print(f"Manufactured solution: u_r = {a:.1e}·r, u_z = {b:.1e}·z")
print("-" * 70)
print(f"Maximum displacement error: {max_err:.3e}")
print("=" * 70 + "\n")
return out